Submitting data and metadata to the TaRGET II DCC¶
Log in to the Submission Interface (https://submit.targetepigenomics.org)
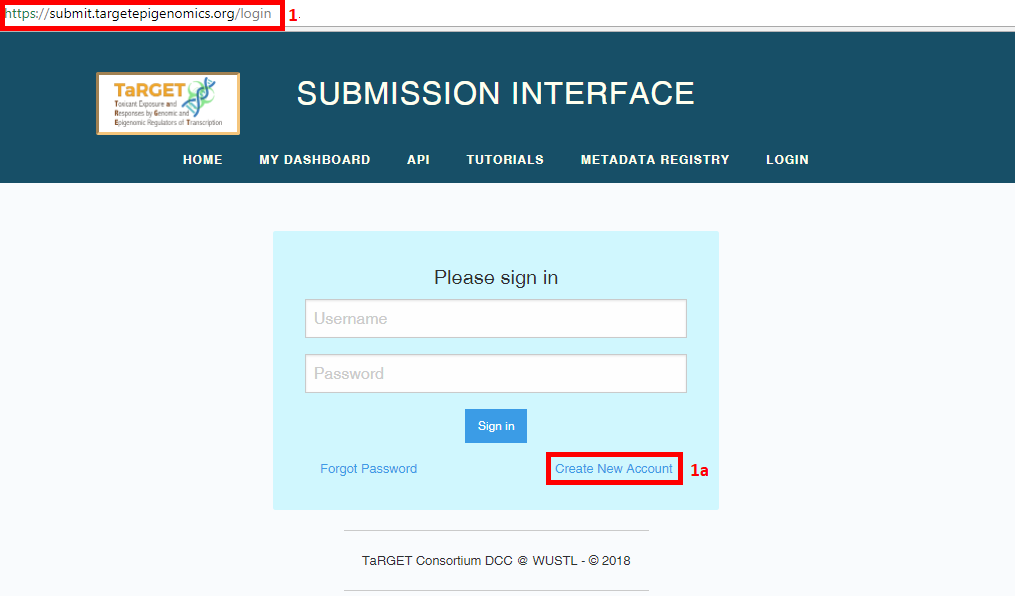
- On your first login, you will need to register as a new user (“Create New Account”). After completing all fields, select “Create”, then “Login”. If you have a consortium wiki account, you should use the same username and email for this account.
Go to your Submission Dashboard (https://submit.targetepigenomics.org/dashboard)
- You can get to your Submission Dashboard by selecting the “View my Submission Dashboard” button on your homepage or selecting “My Dashboard” from the banner.
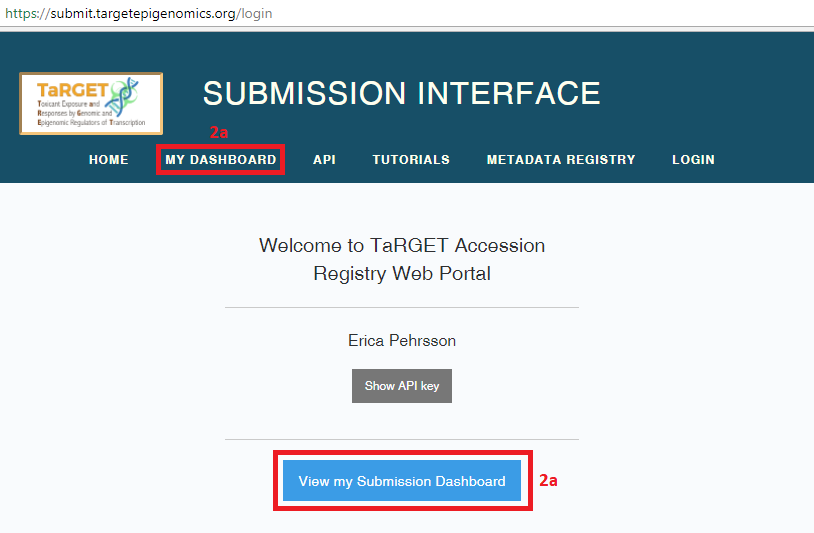
- From here, you may choose to register metadata or data by selecting the appropriate button.
Data submission¶
Create a data submission
- Select the “Data” button on your dashboard. You will be taken to a page (https://submit.targetepigenomics.org/submissions/list) that lists your previous data submissions (if any) and allows you to start a new submission.
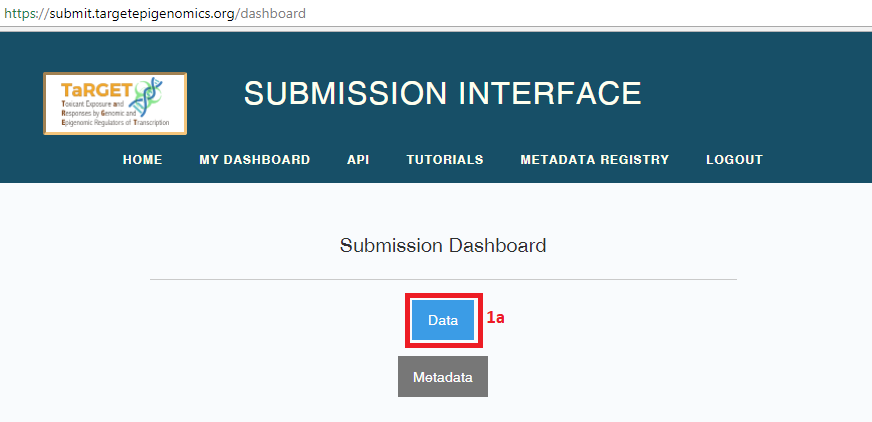
- To create a new submission, select “Start new data file submission - SFTP”. The Aspera upload option has been deprecated.
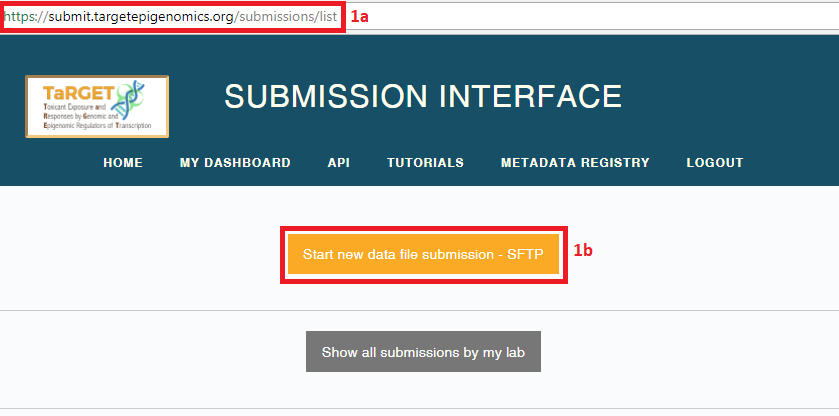
- Specify your lab and username and other basic information for your submission. You may only include files from one assay (e.g., ATAC-seq, RNA-seq) in a single submission, and they must all be single-end or paired-end. Select “Submit”.
- Review your entries and either confirm that they are correct (“Yes”) or return to the form to make changes (“No”).
- Next, follow the instructions to generate an md5sum list for your files, which will be used to ensure that the files were completely uploaded. Please also note the new file naming requirements. Submitted files must be in fastq or fastq.gz format. After pasting the md5sum list in the window, select “Submit” at the bottom of the page. An error will be returned if the naming convention has not been followed or the number of rows in the md5sum list does not match the number of files specified on the previous page.
- For pilot data, you will fill out a limited set of metadata fields during data submission. To fill out metadata, select files in the left pane, then select the relevant metadata for those files in the right pane and hit “Save”. On the review panel, select “Yes” if the metadata is correct. Repeat for all files. Files will be removed from the list once metadata has been registered for them, but you will have the chance to correct all metadata in the next step.
- Select “Review”, edit the table of metadata for each file as needed, and select “Finish”.
- The final page provides detailed instructions for uploading your data via command line SFTP. Please note that for production data, you must still provide metadata (see below).
View previous data submissions
- Previously registered submissions are listed on your data Submission Dashboard. You may toggle between your submissions and your lab’s submissions by selecting the buttons “Show all submissions by my lab”/”Show my submissions”.
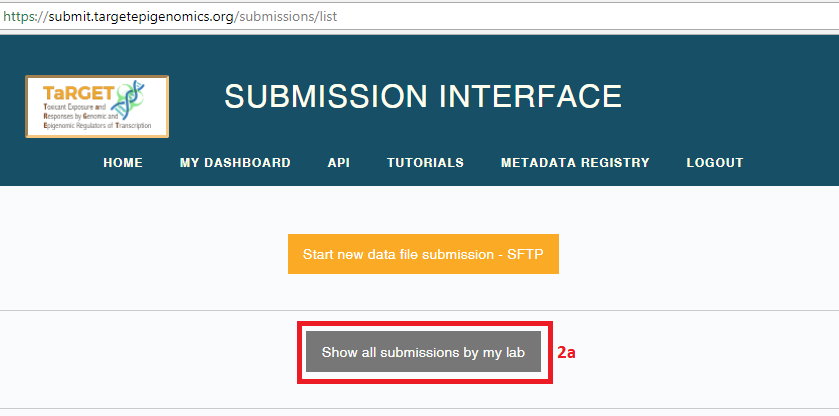
- Each submission is assigned a unique UUID and a DCC data wrangler. The data wrangler is your primary point of contact for the submission.
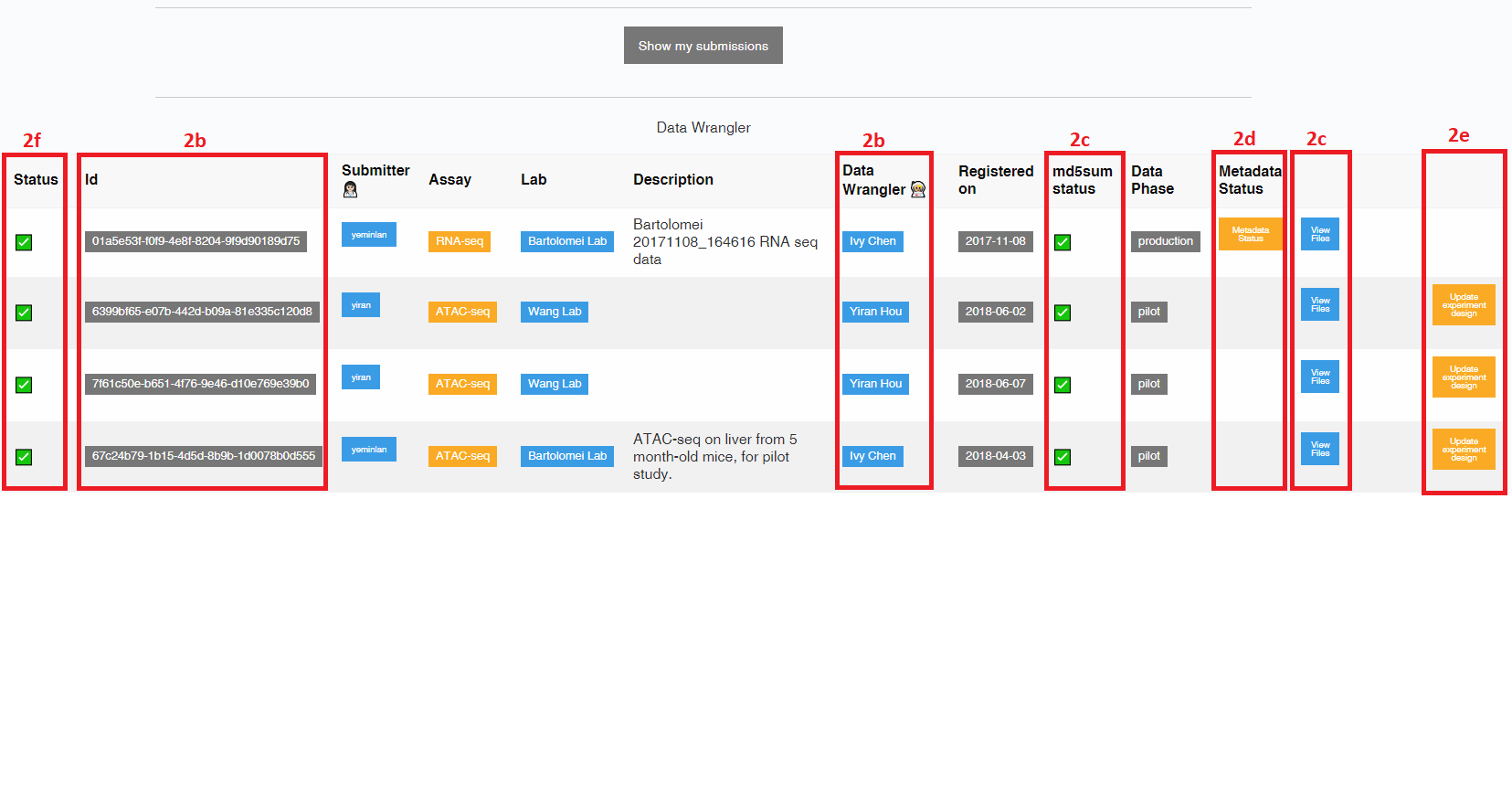
- The md5sum status column shows the results of initial validation performed on your uploaded files. If the number, name, or md5sum of the uploaded files does not match those registered, the status will be a red X and the submission must be corrected before proceeding. To view files that were uploaded as part of a submission, select the “View Files” button. Once files have been uploaded to the DCC server, they will be listed here along with their own UUID and upload date.
- Submissions are not considered complete until metadata has been registered for each file. You can view a report showing the level of completeness of each metadata object attached to each file in the submission by selecting the “Metadata Status” button. Selecting the “Upload Metadata” button will take you to instructions for bulk upload of production metadata (see below).
- Pilot metadata can be updated using the “Update experiment design” button on the submission dashboard - NOT through the Accession Registry.
- Once the submission metadata is complete, your data wrangler will sign off on your submission, and the “Status” column will have a green check. Then, the QC pipeline will be run on the submission, and a QC report will be linked to each file under “View Files”.
Metadata organization¶
For a submission to be marked complete and to be available on the Data Portal, you must fill out several pieces of information about how the files were generated (e.g., how the mouse was treated, how the assay was performed). For production data, complete metadata constitutes all required fields and relationships.
The metadata is organized into discrete categories (such as Mouse, Assay, Reagent) that are linked together. The entity relationship diagram on the Accession Registry Portal home page (https://meta.targetepigenomics.org/) displays the relationships between the metadata categories. Some categories will have only one or a few unique instances per lab (e.g., Bioproject), while others (e.g., “Mouse” for individual mice, “Assay” for experimental assays performed on tissue obtained from a mouse) will have many. By storing metadata as unique objects, we can avoid entering redundant data (e.g., multiple mice may link to the same Diet and Treatment).
The following instructions can be used to register production metadata in the TaRGET II DCC metadata database. You can use them to: 1. Upload new metadata to the database; 2. Update existing records in the database; 3. Establish relationships between metadata records. You can register metadata one-by-one via the Accession Registry (for Lab, Bioproject, Diet, Treatment, and Reagent) or in bulk via the web UI (for Litter, Mouse, Biosample, Assay, and File). Bulk upload via the command line can be performed by request. Please note that pilot metadata should be updated only through the “Update experiment design” button on the submission dashboard.
Metadata submission via the Accession Registry¶
To register metadata one-by-one, go to the Accession Registry Portal (https://meta.targetepigenomics.org/).
- Fill out metadata for your samples by clicking on the metadata objects listed under “Other Metadata”.
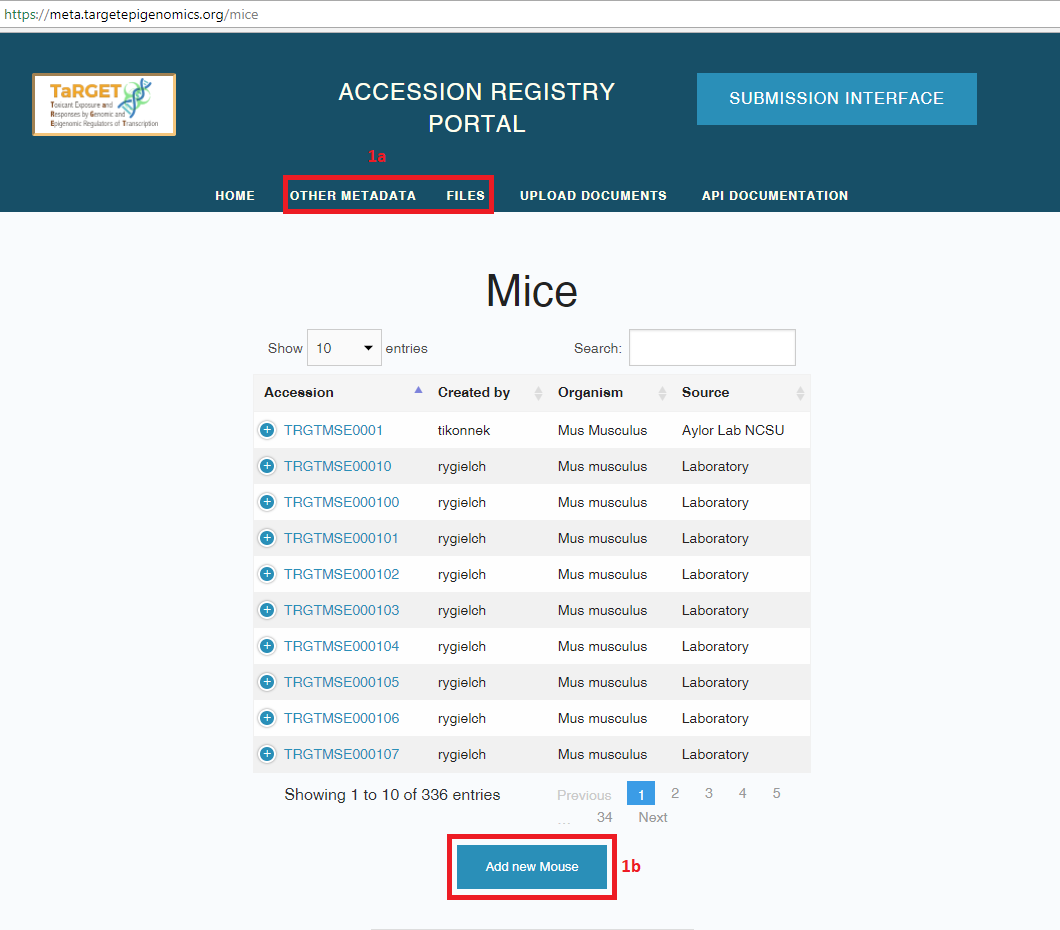
- To create a new metadata object, fill out all of the required fields under the “Add new __” button.
- Some fields will include a description or a drop-down menu of available terms. After you submit the object (“Create”), a notification will appear that the object was successfully created, and its randomly generated, permanent accession number will become available in the list of current objects.
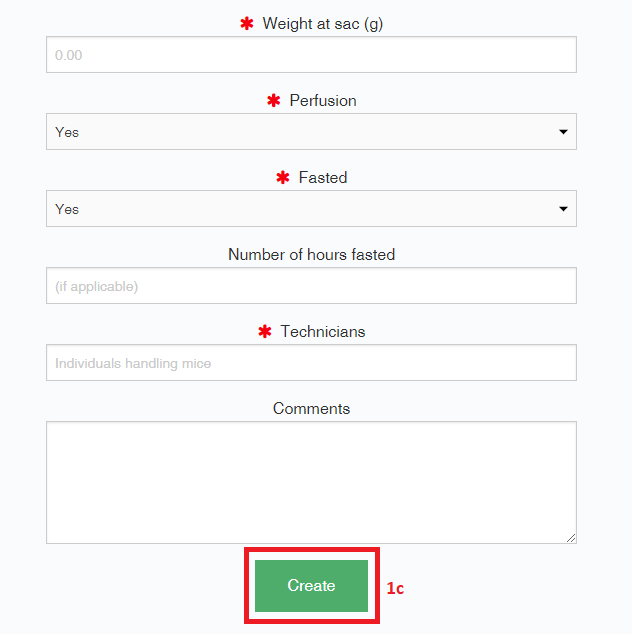
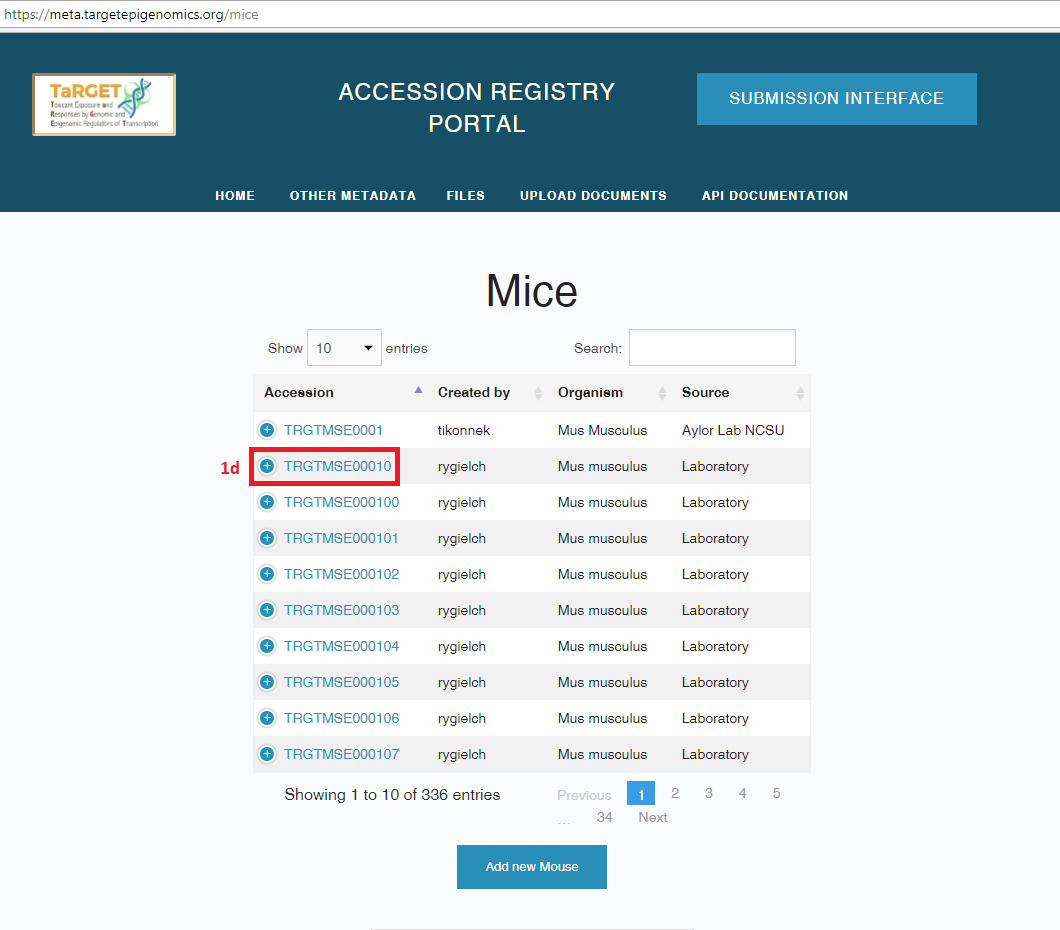
- To view the details of a metadata object, select the accession number for that object. If an object has already been registered, you should not need to register it again when you submit new metadata; however, you should check to make sure that all of the fields match your requirements.
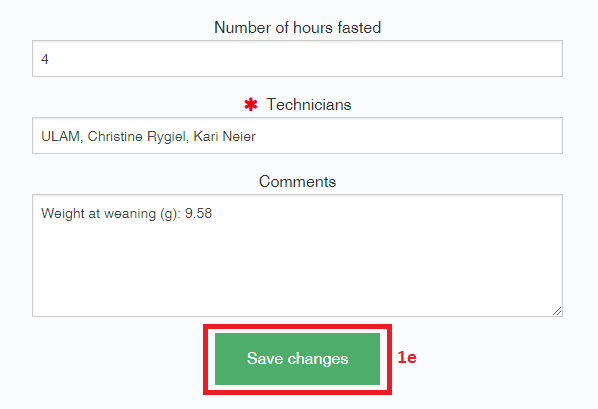
- To edit a metadata object, alter the relevant fields, and select “Save changes”.
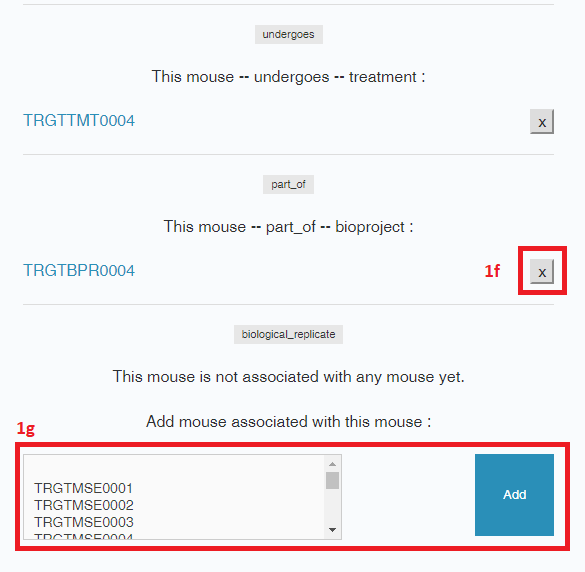
- To link a metadata object to another metadata object (e.g., associate a Bioproject with its Lab), select the object from the drop down menu and select “Add”. Links can also be deleted without deleting the object (“X”).
- To delete a metadata object, select “Delete _”. All links between the object and other objects must be deleted before the object can be deleted.
Metadata bulk submission via the web UI¶
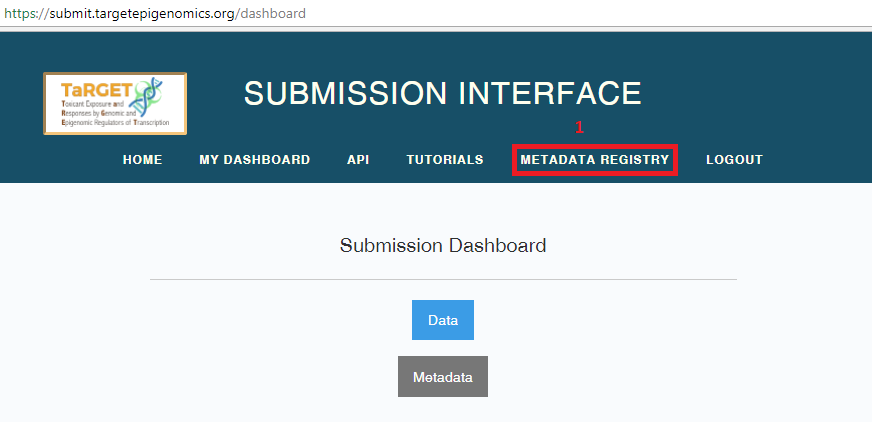
To register metadata in bulk, on your Submission Dashboard (https://submit.targetepigenomics.org/dashboard), select “Metadata” to go to your metadata submission dashboard.
Select “Create/Update bulk metadata submission” to access the web UI for bulk upload of metadata.
Login and follow the tutorial available on this site to download a new blank template or re-download previously submitted metadata into the most recent metadata version.
Instructions for filling out the Excel template:
- All required fields (red headers) and relationships (blue headers) must be populated.
- Many fields are restricted to a list of pre-determined values available in a drop-down menu. To add additional values to the list, contact the Metadata WG.
- Link metadata entries together by entering the relevant System Accession in the blue/teal relationship columns. Metadata can be linked to other records already in the metadata database using their System Accession.
- Enter dates as a string with format “YYYY-MM-DD”.
- All mouse ages should be specified in weeks.
- Please use the following missing data fields: Text, “NA”; Number, “999999999”; Date, “1970-01-01”
- All protocol fields should be linked to finalized protocols that are available on the wiki or the metadata registry, including lab-specific protocols.
Thank you for using the TaRGET DCC submission pipeline! Please contact us with any questions.